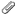Hi ptaylor. I add the option "-giant_move" to my afni_proc.py command and run the afni_proc.py command as you suggest above in the last replay. then i underlayed the final anatomic data and overlayed the error time series "errts.s01.fanaticor" like shown in the picture attached below. i applied a thershhold value to the error time series. is my preprocessing step is alright? because i see correlation in every where except the brain!. am i alright to continue to the next steps?
and i sent a copy of the resulted APQC HTML to the e-mail
i also attached the output of afni_system_check.py -check_all
thank yot ptaylor.