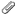Ok, this *seemed* to work and the results look as I'd expect. However, I had to use a -GOFORIT 4 to get it to run, which makes me nervous so I'd like to confirm with you.
The design matrix includes:
columns 0 - 23 - polort
columns 24 - 35 - motion and derivatives
columns 36 - 47 - impulse response matrix as formatted in my previous post
Relavant columns for my example:
36 - 38 are columns for baseline trials
42 - 47 are columns where participants hear a sound
I then created a -glt by making one row with columns 36-38 '-1' and columns 42-47 '1' so that I can create a sound>no sound contrast (all other columns are '0').
Here is my 3dDeconvolve command:
3dDeconvolve \
-input $runs -jobs 24 \
-polort A \
-ortvec ${s}_MoPar_demean.1D demean \
-ortvec ${s}_MoPar_derv.1D derv \
-ortvec ${s}_FIR.1D ir \
-censor ${s}_censor.1D \
-num_glt 1 \
-glt 1 sound-nosound.1D -glt_label 1 s-bl \
-overwrite -bout -tout \
-x1D ${s}.GLM.xmat.1D -x1D_uncensored ${s}.uncensor.GLM.xmat.1D \
-bucket ${s}.GLM.stats -GOFORIT 4
Here is the 3dDeconvolve output:
++ 3dDeconvolve extending num_stimts from 0 to 24 due to -ortvec
++ 3dDeconvolve: AFNI version=AFNI_16.3.20 (Dec 31 2016) [64-bit]
++ Authored by: B. Douglas Ward, et al.
++ current memory malloc-ated = 666,928 bytes (about 667 thousand [kilo])
++ loading dataset S10_V180_1.smooth+tlrc.BRIK S10_V180_2.smooth+tlrc.BRIK S10_V180_3.smooth+tlrc.BRIK S10_V180_4.smooth+tlrc.BRIK S10_V180_5.smooth+tlrc.BRIK S10_V180_6.smooth+tlrc.BRIK
++ current memory malloc-ated = 1,174,677,253 bytes (about 1.2 billion [giga])
++ Auto-catenated input datasets treated as multiple imaging runs
++ Auto-catenated datasets start at: 0 180 360 540 720 900
++ STAT automask has 205353 voxels (out of 271633 = 75.6%)
++ Skipping check for initial transients
++ Imaging duration=360.0 s; Automatic polort=3
++ Number of time points: 1080 (no censoring)
+ Number of parameters: 48 [48 baseline ; 0 signal]
++ total shared memory needed = 107,566,668 bytes (about 108 million [mega])
++ current memory malloc-ated = 1,174,966,124 bytes (about 1.2 billion [giga])
++ mmap() memory allocated: 107,566,668 bytes (about 108 million [mega])
++ Memory required for output bricks = 107,566,668 bytes (about 108 million [mega])
++ Wrote matrix values to file S10.GLM.xmat.1D
++ ========= Things you can do with the matrix file =========
++ (a) Linear regression with ARMA(1,1) modeling of serial correlation:
3dREMLfit -matrix S10.GLM.xmat.1D \
-input "S10_V180_1.smooth+tlrc.BRIK S10_V180_2.smooth+tlrc.BRIK S10_V180_3.smooth+tlrc.BRIK S10_V180_4.smooth+tlrc.BRIK S10_V180_5.smooth+tlrc.BRIK S10_V180_6.smooth+tlrc.BRIK" \
-tout -Rbuck S10.GLM.stats_REML -Rvar S10.GLM.stats_REMLvar -verb
++ N.B.: 3dREMLfit command above written to file S10.REML_cmd
++ (b) Visualization/analysis of the matrix via ExamineXmat.R
++ (c) Synthesis of sub-model datasets using 3dSynthesize
++ ==========================================================
++ Wrote matrix values to file S10.uncensor.GLM.xmat.1D
++ ----- Signal+Baseline matrix condition [X] (1080x48): 3.84811 ++ VERY GOOD ++
*+ WARNING: !! in Signal+Baseline matrix:
* Largest singular value=1.86247
* 1 singular value is less than cutoff=1.86247e-07
* Implies strong collinearity in the matrix columns!
++ Signal+Baseline matrix singular values:
0 0.125775 0.152245 0.186387 0.210547
0.297469 0.315213 0.524212 0.563959 0.710978
0.80135 0.908348 0.927272 0.953512 0.969709
0.982664 0.98997 0.995209 0.997447 0.999185
0.99998 1 1 1.00023 1.00086
1.00154 1.00536 1.00797 1.0121 1.01273
1.01361 1.02203 1.03139 1.03458 1.0483
1.06934 1.07048 1.07709 1.08865 1.09839
1.12271 1.14064 1.22223 1.27599 1.42174
1.52205 1.66472 1.86247
++ ----- Baseline-only matrix condition [X] (1080x48): 3.84811 ++ VERY GOOD ++
*+ WARNING: !! in Baseline-only matrix:
* Largest singular value=1.86247
* 1 singular value is less than cutoff=1.86247e-07
* Implies strong collinearity in the matrix columns!
++ Baseline-only matrix singular values:
0 0.125775 0.152245 0.186387 0.210547
0.297469 0.315213 0.524212 0.563959 0.710978
0.80135 0.908348 0.927272 0.953512 0.969709
0.982664 0.98997 0.995209 0.997447 0.999185
0.99998 1 1 1.00023 1.00086
1.00154 1.00536 1.00797 1.0121 1.01273
1.01361 1.02203 1.03139 1.03458 1.0483
1.06934 1.07048 1.07709 1.08865 1.09839
1.12271 1.14064 1.22223 1.27599 1.42174
1.52205 1.66472 1.86247
++ ----- stim_base-only matrix condition [X] (1080x24): 2.28456 ++ VERY GOOD ++
*+ WARNING: !! in stim_base-only matrix:
* Largest singular value=1.60031
* 1 singular value is less than cutoff=1.60031e-07
* Implies strong collinearity in the matrix columns!
++ stim_base-only matrix singular values:
0 0.306619 0.324891 0.474359 0.558376
0.600785 0.730422 0.826685 0.886871 0.987358
1.00504 1.01798 1.02599 1.02889 1.03712
1.06551 1.0745 1.0774 1.07927 1.13786
1.37112 1.41741 1.52672 1.60031
++ ----- polort-only matrix condition [X] (1080x24): 1.01269 ++ VERY GOOD ++
*+ WARNING: +++++ !! Matrix inverse average error = 0.00520833 ** BEWARE **
++ Matrix setup time = 1.10 s
*+ WARNING: !! 3dDeconvolve -GOFORIT is set to 4: running despite 4 matrix warnings
*+ WARNING: !! See file 3dDeconvolve.err for all WARNING and ERROR messages !!
*+ WARNING: !! Please be sure you understand what you are doing !!
*+ WARNING: !! If in doubt, consult with someone or with the AFNI message board !!
Please let me know if any of those warnings are actually concerning. I have a hunch that the warnings are because it thinks that all -ortvec parameters will go into baseline. There are no collinearity issues in my impulse response regressors. The range of collinearity between the regressors I'm interested in is r = [-0.15 -0.03].
Thanks!
Dustin