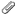Hm, so the alignment trims off part of the T1 and it seems to be related to square surrounding the dwi, thus limiting the area (photo attached)
fat_proc_align_anat_pair -in_t1w $SUBJECTS_DIR/$subj/mri/T1.nii.gz -in_t2w $dtipath/$subj/data.nii.gz -out_t2w_grid -prefix t1_aligned
I'm not sure if the output is informative but I am pasting it here:
++ Start script version: 1.2
++ Found input file: /home/eji/projects/spins/preproc/freesurfer/subjects/w0/SPN01_CMH_0001_01/mri/T1.nii.gz
++ Found input file: /home/eji/data3/spins/dwi/SPN01_CMH_0001_01/data.nii.gz
++ Based on prefix, the output directory will be:
.
++ Based on prefix, the output prefix will be:
t1_aligned
++ Making working directory: ./__WORKING_align_anat_pair_t1_aligned
++ Note: final volume grid resolution will be 1.0 mm isotropic.
-----> STARTING fat_proc_align_anat_pair ---->
This command:
fat_proc_align_anat_pair -in_t1w /home/eji/projects/spins/preproc/freesurfer/subjects/w0/SPN01_CMH_0001_01/mri/T1.nii.gz -in_t2w /home/eji/data3/spins/dwi/SPN01_CMH_0001_01/data.nii.gz -out_t2w_grid -prefix t1_aligned
++ Echoing the command to: ./t1_aligned_cmd.txt
++ Unifize T1w proc vol.
++ 3dUnifize: AFNI version=AFNI_20.1.06 (May 4 2020) [64-bit]
+ Pre-processing: ADV...........UW[s192914]Gm
++ Output dataset ./__WORKING_align_anat_pair_t1_aligned/f01_t1uni.nii
++ ===== Elapsed = 26.5 sec
++ Threshold T1w proc vol.
1281.796509
++ 3dcalc: AFNI version=AFNI_20.1.06 (May 4 2020) [64-bit]
++ Authored by: A cast of thousands
++ Output dataset ././__WORKING_align_anat_pair_t1_aligned/f02_t1thr.nii
++ Calculating alignment (t2w -> t1w) matrix.
+++ Command Echo:
3dAllineate -1Dmatrix_save ./__WORKING_align_anat_pair_t1_aligned/map_t2w_to_t1w.aff12.1D -prefix ./__WORKING_align_anat_pair_t1_aligned/f05_t1align.nii -source /home/eji/data3/spins/dwi/SPN01_CMH_0001_01/data.nii.gz -base ./__WORKING_align_anat_pair_t1_aligned/f02_t1thr.nii -twopass -cost lpc -cmass -warp shift_rotate -source_automask -wtprefix ./__WORKING_align_anat_pair_t1_aligned/wtvol.nii.gz -autoweight -final wsinc5 -overwrite
++ 3dAllineate: AFNI version=AFNI_20.1.06 (May 4 2020) [64-bit]
++ Authored by: Zhark the Registrator
++ Source dataset: /home/eji/data3/spins/dwi/SPN01_CMH_0001_01/data.nii.gz
++ Base dataset: ./__WORKING_align_anat_pair_t1_aligned/f02_t1thr.nii
++ Loading datasets into memory
++ Output dataset ./__WORKING_align_anat_pair_t1_aligned/wtvol.nii.gz
++ Local correlation: blok type = 'RHDD(6.54321)'
++ master dataset for output = base
++ OpenMP thread count = 15
++ ======= Allineation of 65 sub-bricks using Local Pearson Correlation Signed =======
volume 0++ *** Coarse pass begins ***
++ *** Fine pass begins ***
+ Final fine fit Parameters:
x-shift= -1.2558 y-shift= 20.3702 z-shift= 12.0026 ... enorm= 23.6766 mm
z-angle= 0.3099 x-angle= 1.1459 y-angle= 5.0282 ... total= 5.1693 deg
++ wsinc5 interpolation setup:
+ taper function = Min sidelobe 3 term
+ taper cut point = 0.000
+ window radius = 5 voxels
+ window shape = Cubical
+ The above can be altered via the AFNI_WSINC5_* environment variables.
+ (To avoid this message, 'setenv AFNI_WSINC5_SILENT YES'.)
+ wsinc5 CUBE(5) mask has 1000 points
..1++ *** Coarse pass begins ***
..2++ *** Coarse pass begins ***
..3++ *** Coarse pass begins ***
..4++ *** Coarse pass begins ***
..5++ *** Coarse pass begins ***
..6++ *** Coarse pass begins ***
..7++ *** Coarse pass begins ***
..8++ *** Coarse pass begins ***
..9++ *** Coarse pass begins ***
..10++ *** Coarse pass begins ***
..11++ *** Coarse pass begins ***
..12++ *** Coarse pass begins ***
..13++ *** Coarse pass begins ***
..14++ *** Coarse pass begins ***
..15++ *** Coarse pass begins ***
..16++ *** Coarse pass begins ***
..17++ *** Coarse pass begins ***
..18++ *** Coarse pass begins ***
..19++ *** Coarse pass begins ***
..20++ *** Coarse pass begins ***
..21++ *** Coarse pass begins ***
..22++ *** Coarse pass begins ***
..23++ *** Coarse pass begins ***
..24++ *** Coarse pass begins ***
..25++ *** Coarse pass begins ***
..26++ *** Coarse pass begins ***
..27++ *** Coarse pass begins ***
..28++ *** Coarse pass begins ***
..29++ *** Coarse pass begins ***
..30++ *** Coarse pass begins ***
..31++ *** Coarse pass begins ***
..32++ *** Coarse pass begins ***
..33++ *** Coarse pass begins ***
..34++ *** Coarse pass begins ***
..35++ *** Coarse pass begins ***
..36++ *** Coarse pass begins ***
..37++ *** Coarse pass begins ***
..38++ *** Coarse pass begins ***
..39++ *** Coarse pass begins ***
..40++ *** Coarse pass begins ***
..41++ *** Coarse pass begins ***
..42++ *** Coarse pass begins ***
..43++ *** Coarse pass begins ***
..44++ *** Coarse pass begins ***
..45++ *** Coarse pass begins ***
..46++ *** Coarse pass begins ***
..47++ *** Coarse pass begins ***
..48++ *** Coarse pass begins ***
..49++ *** Coarse pass begins ***
..50++ *** Coarse pass begins ***
..51++ *** Coarse pass begins ***
..52++ *** Coarse pass begins ***
..53++ *** Coarse pass begins ***
..54++ *** Coarse pass begins ***
..55++ *** Coarse pass begins ***
..56++ *** Coarse pass begins ***
..57++ *** Coarse pass begins ***
..58++ *** Coarse pass begins ***
..59++ *** Coarse pass begins ***
..60++ *** Coarse pass begins ***
..61++ *** Coarse pass begins ***
..62++ *** Coarse pass begins ***
..63++ *** Coarse pass begins ***
..64
++ *** Coarse pass begins ***
++ Output dataset ./__WORKING_align_anat_pair_t1_aligned/f05_t1align.nii
++ Wrote -1Dmatrix_save ./__WORKING_align_anat_pair_t1_aligned/map_t2w_to_t1w.aff12.1D
++ 3dAllineate: total CPU time = 0.0 sec Elapsed = 1848.3
++ ###########################################################
++ # Please check results visually for alignment quality #
++ ###########################################################
++ Inverting alignment matrix (to get t1w -> t2w).
++ Apply alignment matrix.
++ Outputting final image to t2w space
+++ Command Echo:
3dAllineate -1Dmatrix_apply ./__WORKING_align_anat_pair_t1_aligned/map_t1w_to_t2w.aff12.1D -source ./__WORKING_align_anat_pair_t1_aligned/f00_t1res.nii -master /home/eji/data3/spins/dwi/SPN01_CMH_0001_01/data.nii.gz -prefix ./__WORKING_align_anat_pair_t1_aligned/f06_t1mapped.nii -float -final wsinc5
++ 3dAllineate: AFNI version=AFNI_20.1.06 (May 4 2020) [64-bit]
++ Authored by: Zhark the Registrator
++ Source dataset: ./__WORKING_align_anat_pair_t1_aligned/f00_t1res.nii
++ Base dataset: (not given)
++ Loading datasets into memory
+ -cmass x y z shifts = 0.000 0.000 0.000
+ shift search range is +/- = 81.855 81.855 81.855
++ OpenMP thread count = 15
++ ========== Applying transformation to 1 sub-bricks ==========
++ wsinc5 interpolation setup:
+ taper function = Min sidelobe 3 term
+ taper cut point = 0.000
+ window radius = 5 voxels
+ window shape = Cubical
+ The above can be altered via the AFNI_WSINC5_* environment variables.
+ (To avoid this message, 'setenv AFNI_WSINC5_SILENT YES'.)
+ wsinc5 CUBE(5) mask has 1000 points
++ Output dataset ./__WORKING_align_anat_pair_t1_aligned/f06_t1mapped.nii
++ 3dAllineate: total CPU time = 0.0 sec Elapsed = 1.7
++ ###########################################################
++ Perhaps for prepping for FreeSurfing:
-> making sure that there are even numbers of voxels
along each spatial dimension.
++ The dimensions are: 128 128 71
+* Found an odd dimension in the [3]th axis!
--> will hopefully fix.
++ Zeropadding: -I 1
+++ Command Echo:
3dZeropad -overwrite -prefix ./__WORKING_align_anat_pair_t1_aligned/f08_t1padded.nii -I 1 ./__WORKING_align_anat_pair_t1_aligned/f06_t1mapped.nii
++ 3dZeropad: AFNI version=AFNI_20.1.06 (May 4 2020) [64-bit]
++ output dataset: ./__WORKING_align_anat_pair_t1_aligned/f08_t1padded.nii
++ Done workin'. Just need to copy the final dset.
++ 3dcalc: AFNI version=AFNI_20.1.06 (May 4 2020) [64-bit]
++ Authored by: A cast of thousands
++ Output dataset ././t1_aligned.nii.gz
++ More QC images: b0 on initial ref.
++ 3dedge3: AFNI version=AFNI_20.1.06 (May 4 2020) [64-bit]
++ QC image 00 (./t1_aligned.nii.gz edges on /home/eji/data3/spins/dwi/SPN01_CMH_0001_01/data.nii.gz): t1_aligned_qc_t2w_t1wE
++ My command:
@chauffeur_afni -ulay /home/eji/data3/spins/dwi/SPN01_CMH_0001_01/data.nii.gz -ulay_range 2% 98% -olay ./__WORKING_align_anat_pair_t1_aligned/f10_edges.nii -func_range_perc_nz 50 -pbar_posonly -cbar red_monochrome -opacity 6 -prefix ./QC/t1_aligned_qc_t2w_t1wE -montx 5 -monty 3 -set_xhairs OFF -label_mode 1 -label_size 3 -do_clean
------------------ start of optionizing ------------------
++ Found input file: /home/eji/data3/spins/dwi/SPN01_CMH_0001_01/data.nii.gz
++ Found input file: ./__WORKING_align_anat_pair_t1_aligned/f10_edges.nii
++ Using blowup factor: 2
++ Using opacity: 6
++ Making temporary work directory to copy vis files: ./QC/__tmp_chauf_oC6qIY1FE4T
++ Converted 0 to labels
++ Final subbrick indices: -1 -1 -1
++ Copy ulay to visualize (volumetric) within user's range:
** FATAL ERROR: -percentile can only be used on one sub-brick only.
Use sub-brick selectors '[.]' to specify sub-brick of interest.
** Program compile date = May 4 2020
** FATAL ERROR: -percentile can only be used on one sub-brick only.
Use sub-brick selectors '[.]' to specify sub-brick of interest.
** Program compile date = May 4 2020
amin: Subscript out of range.
++ QC image 01 (./t1_aligned.nii.gz olay on /home/eji/data3/spins/dwi/SPN01_CMH_0001_01/data.nii.gz): t1_aligned_qc_t2w_t1w
++ My command:
@chauffeur_afni -ulay /home/eji/data3/spins/dwi/SPN01_CMH_0001_01/data.nii.gz -ulay_range 2% 98% -olay ./t1_aligned.nii.gz -pbar_posonly -opacity 4 -prefix ./QC/t1_aligned_qc_t2w_t1w -montx 5 -monty 3 -set_xhairs OFF -label_mode 1 -label_size 3 -do_clean
------------------ start of optionizing ------------------
++ Found input file: /home/eji/data3/spins/dwi/SPN01_CMH_0001_01/data.nii.gz
++ Found input file: ./t1_aligned.nii.gz
++ Using blowup factor: 2
++ Using opacity: 4
++ Making temporary work directory to copy vis files: ./QC/__tmp_chauf_xhg9XLwFz8m
++ Converted 0 to labels
++ Final subbrick indices: -1 -1 -1
++ Copy ulay to visualize (volumetric) within user's range:
** FATAL ERROR: -percentile can only be used on one sub-brick only.
Use sub-brick selectors '[.]' to specify sub-brick of interest.
** Program compile date = May 4 2020
** FATAL ERROR: -percentile can only be used on one sub-brick only.
Use sub-brick selectors '[.]' to specify sub-brick of interest.
** Program compile date = May 4 2020
amin: Subscript out of range.
++ Cleaning working directory!
++ The final data set is here: ./t1_aligned.nii.gz
++ The final spatial dims are: 128 128 72
++ The spatial resolution is: 2.000000 2.000000 2.000000