2. Starting with AFNI¶
2.1. Overview¶
Welcome to the AFNI software toolbox. AFNI (Analysis of Functional NeuroImages) is a suite of programs for looking at and analyzing MRI brain images at all stages of analysis (planning, setting up acquisiton, preprocessing, analysis, quality control and statistical analysis). It contains C, Python and R programs, as well as shell scripts.
The philosophy driving AFNI development is simple: help keep users close to their data.
AFNI is primarily developed for the processing, analysis and display of multiple MRI modalities:
anatomical/structural MRI
at various field strengths
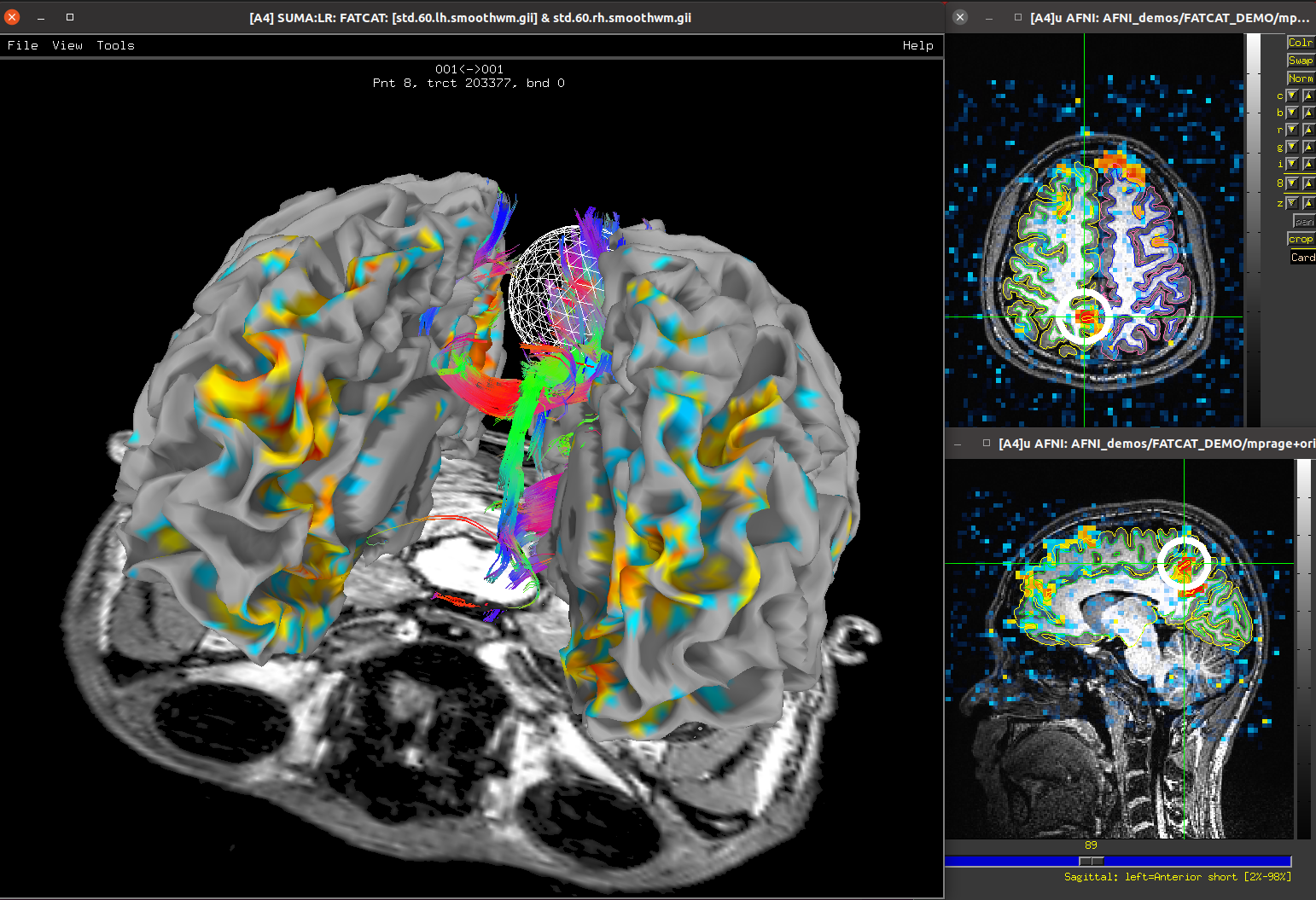
diffusion weighted imaging (DWI)
for DTI or HARDI modeling and tractography (using the FATCAT programs)
functional MRI (FMRI)
resting state, task-based or naturalistic paradigms
single- or multi-echo acquisitions
voxelwise or ROI-based analyses, volumetric or surface-based
convenient+efficient pipeline design with
afni_proc.py
Many AFNI programs have also been applied and adapted to other modalities, such as ECoG, EEG, MEG, and more.
AFNI can be used for volumetric or surface analyses, as well as their combination, and has been applied widely across studies of humans, NHP and other species.
2.2. Where to begin¶
Below we provide a quick roadmap through the AFNI toolbox, including educational resources, reference links, demo examples (with data+scripts) and more. Note: these are just subsets of the material here, which we encourage exploring.
Installation instructions
Bootcamp and video lectures
AFNI Academy series: lectures based on the AFNI Bootcamp, combining theory and hands-on examples
Links to AFNI handouts (part of Bootcamp), GUI keyboard shortcuts and startup tips (and more)
Program listings and helps
Code, command, script and functionality examples
Additional resources
Code-centric
2.3. Asking questions and following updates¶
We are happy to answer questions, receive suggestions, and generally discuss data analysis on the AFNI Message Board.
Additionally, please consider signing up for the AFNI Digest. This approximately weekly (or so) emailer contains developments or changes within the AFNI realm that we deem large enough for wide dissemination. We will also broadcast things like Bootcamp info; installation and build instruction changes; processing commentary; and perhaps dollops of humor.
