7.5. 3dTrackID: making tracts¶
7.5.1. Overview¶
There are a few modes/varieties/flavors of fiber tracking which can be
performed with 3dTrackID. Selecting which one to use depends a
bit on the available inputs (though, these are mostly the same for
each mode) and on the desired outcome (“with great power comes great
responsibility”).
A general theme of all of them is to be able to deal with the following scenario: I have a network of target ROIs; where is the most likely location of the white matter connecting any of them, and what quantitative properties do those white matter ROIs have?
Excited? Me, too. Before continuing, though, let’s agree on some terminology with which these help pages will hopefully show consistency:
The set of things among which we want to find connections are referred to as target ROIs or just targets, and collectively as the network of targets. There are usually N>1 targets in a network, but sometimes it is useful to use a wholebrain mask as a ‘network of one’ for viewing tracts. Each target is a set of voxels of a particular nonzero integer, and we may refer to it either by the name of the integer or by a label associated with that integer (e.g., “1” is the “precuneus”, etc.). The use of
3dROIMakerin forming target networks is described in 3dROIMaker: making (networks of) ROIs.The set of tracts connecting a given pair of targets is called a bundle. These are specifically the collection of linear structures themselves. Heck, sometimes we’ll abandon all formality and just call them ‘tracts’, as well.
The volume containing a particular bundle is called a WM ROI. This refers to the set of voxels containing the particular tracts of interest between targets. We try to keep the terminology distinct from the set of target ROIs to avoid confusion.
Bundles and WM ROIs can be made use AND-logic and/or OR-logic (enough conjunctions in that statement for you?):
AND-logic means connecting two distinct targets, such as (target) ROI 4 and ROI 5; in output files, these types of connections are grouped under the name PAIR or PAIRMAP (for “pairwise” target bundles). NB: a PAIR tract could pass through a third target as well, but it doesn’t matter at all for labeling or its definition as an AND-logic connection.
OR-logic means passing through at least one specific target, such as target ROI 2, with the possibility (but not the necessity) of passing through any other target(s); in output files, these types of connections are grouped under the name INDI or INDIMAP (for ‘individual’ target bundles). NB: for purely notational and labelling purposes, it’s also convenient to think of an INDI bundle being the special case of connecting a target with itself– this is useful for the matrix-style storing of information described in the next list item.
the PAIR tracts/bundles and WM ROIs are a subset of INDI ones.
Tracking to determine the structural connectivity among pairs/individual targets is analogous to using correlation to determine functional connectivity among a (target) network. Functional connectivity information is generally stored and viewed as correlation matrices, and we borrow the same to store
3dTrackIDoutput in structural connectivity matrices:a given matrix can store properties such as the number of tracts between two targets, the average FA of a WM ROI, etc.
just as in a correlation matrix, each row and column is labeled by a particular target. Each element of the matrix contains the property of the WM ROI that connects those targets.
a diagonal matrix element, where the row number is the same as the wcolumn number, describes a property of an OR-logic (or INDI) WM ROI.
an off-diagonal element describes a property of an AND-logic (or PAIR) WM ROI.
the structural connectivity matrix is symmetric: the property between targets 1 and 7 is the same as that between targets 7 and 1 (and similarly for any off-diagonal element). This is another way of stating that there is no directionality between the structural connectivity properties.
Note
“All targets are equal,” meaning that tracking is performed amongst and within the network of targets, with no special preference. Other programs may have an approach of tracking ‘to’ a target ‘from’ a seed, but this directional approach (where “some targets are more equal than others”) is not the philosphical choice here. Why should a tract result connecting targets 1 and 9 be different than that connecting 9 and 1?
7.5.2. Modus operandi¶
Preface. Once upon a time, there were two separate FATCAT programs for performing deterministic and probabilistic tractography (3dTrackID and 3dProbTrackID, respectively), as described in the initial paper (Taylor and Saad, 2013). Soon after this introduction, it became apparent that there could be only one. After a brief quickening, 3dTrackID picked up the probabilistic attributes and now that program is run in separate modes. Which brings us to the present.
- Current times. There are three distinct modes for performing
tractography, and each is denoted with a short string after the required switch
-mode *. These are:deterministic (
DET),mini-probabilistic (
MINIP),(full) probabilistic (
PROB).
Thus, all FATCAT tracking commands will start with a selection of one of these modes:
3dTrackID -mode {DET|MINIP|PROB} ...
There is a lot of overlap in what kind of data are input and output for these modes. First, we describe the default and optional outputs of all; then, special outputs of some; finally, the differences in inputs (and why they exist as such).
The outputs can be viewed variously and interactively in AFNI and SUMA
(such as for volume, tract, and dset files). Additionally, matrices
of properties can be viewed and saved from the command line with some
fat_*.py functions. Finally, outputs can be used for quantitative
comparison and statistical modeling– one method for doing the latter
exists using G. Chen’s 3dMVM (see below for some description, and the
FATMVM demo introduced Demo data sets and scripts).
7.5.3. Outputs common to all modes¶
By default, each of the
3dTrackIDmodes will output the following:volumes of WM ROIs, both a single PAIRMAP file of the AND-logic connections and a single INDIMAP file of the OR-logic ones. These can be viewed most easily using the AFNI viewer to get a visualization of:
all the locations where tracts went through the network ([0]th brick of either MAP file);
all the locations where tracts went through an individual target ([i]th brick of either MAP file, where i>0);
a grid file (ending with
*.grid), which contains all the structural connectivity matrices for the given network. Quantities include both mean and standard deviation of DTI parameters (FA, MD, L1 and RD), as well as the volume of the WM ROIs (both in terms of physical units, number of voxels, and volume scaled by whole brain mask), as well as the number of tracts. The matrices in these files can be:selected, viewed and saved to an image file using
fat_mat_sel.py;used for group-based statistics with G. Chen’s 3dMVM program, with some helper
fat_mvm*.pyfunctions available for putting everything together and building commands+models.
a dset file (ending with
*.dset), which also contains all of the structural connectivity matrices for a given network. Matrices in these files can be:loaded into SUMA (
$ suma -gdset NAME.niml.dset ...);viewed in SUMA as either a standard, colorful matrix, or as a graph-like network of nodes and edges throughout the 3D brain representation;
Example 1. For example, running
3dTrackIDwith-prefix o.NETS(and-nifti) will produce the output files:o.NETS_000.grid o.NETS_000.niml.dset o.NETS_000_INDIMAP.nii.gz o.NETS_000_PAIRMAP.nii.gz
Comments on these outputs:
A PAIRMAP is not output if the input network has only one target ROI, such as if one is doing a simple whole brain tracking.
One can turn off INDIMAP and PAIRMAP output altogether, using the switch
-no_indipair_out. This might be useful if you are tracking through a large network of targets (for example, something connectome-y) and don’t want to risk having a single reaaally big output file wasting space or causing trouble.By default, all volumetric outputs (PAIRMAP, INDIMAP,
-dump_rois *files, etc.) are in BRIK/HEAD file format. If you prefer NIFTI, you can use the switch-niftito get all “*.nii.gz” files.
Additionally, each mode can also output:
a set of maps/masks of each individual WM ROI. This is done using the option
-dump_rois {AFNI|DUMP|BOTH|AFNI_MAP}. The keyword options each produces a set of individual files of the following:DUMP->3dmaskdump-like text files of each WM ROI (which could take quite a lot of space and not be so useful;AFNI-> binary masks of each WM ROI;BOTH-> both the binary masks and text files (combined outputs ofDUMPandAFNI; the name reflects that it was developed when there were only two individual output formats);AFNI_MAP–> non-binarized maps of each WM ROI, where the value of each voxel is the number of tracts that went through it for that given connection;
Example 1 (continued). Additionally, if one also included the command
-dump_rois AFNI, then the output would include a directory o.NETS/ with the following files, such as:NET_000_ROI_001_001.nii.gz NET_000_ROI_001_004.nii.gz NET_000_ROI_002_002.nii.gz NET_000_ROI_002_003.nii.gz NET_000_ROI_002_004.nii.gz NET_000_ROI_002_006.nii.gz NET_000_ROI_002_007.nii.gz ...
With the specific dump option used here, each file would contain a binary mask of the given WM connection. The file naming convention is: NET_X_ROI_Y_Z.nii.gz, where:
‘X’ is the number of the network (because multiple ones can be tracked simultaneously
‘Y’ is the number or label of a target ROI
‘Z’ is the number or label of another target ROI
The files where ‘Y’==’Z’ contain INDIMAP information of a target, and the others where not(‘Y’==’Z’) contain PAIRMAPs. It’s important to note that tracts will not be found between every possible pair of targets, and so not every possible pairwise combination will have a file output. |
Note
Probably using one of the options
-dump_rois {AFNI|AFNI_MAP}would be the most useful. Some unnamed user(s) would even go so far as to recommend using it all the time, because either would provide the only unambiguous maps of individual WM ROIs output by3dTrackID.A labeltable file (
*.niml.lt) will also be output if one has been attached to the input network file. While one might not view this on its own, having a labeltable set up can be very useful, for example in helping to discuss specific bundles by the anatomical locations they connect.
7.5.4. Outputs specific to {DET|MINIP} modes¶
The outputs in the previous section are output for all modes of
3dTrackID. However, careful readers will note that none of those tractographic outputs actually contained the tracts themselves! These are only output in{DET|MINIP}modes, as the following:a tract file (ending with
*.tract), which contains all the individual tract sequences. Additionally, it internally has the tracts organized into sets of bundles between targets, so that each bundle could be displayed as a separate color. These files are viewable in SUMA, loading with:suma -tract PREFIX.niml.tract ...
One can also load in the dset simultaneously and view the connectivity matrix elements as coloration of tract bundles, such as after:
suma -tract PREFIX.niml.tract -gdset PREFIX.niml.dset ...
(In fact, the dset loaded in could be either one output by
3dTrackIDor by3dNetCorr.)a TRK-format file,
*.trk, legacy of when tractographic output had to be viewed with non-AFNI/SUMA options, which in this case were with TrackVis. These are not output by default. To have these be output, use the the-do_trk_outswitch.
When outputting tract files, one has to choose whether to use AND-logic or OR-logic within the network. That is, whether to keep tracts that have a minimal requirement of going through one target (OR), or whether to require at tract to connect at least two targets (AND). The choice is made using the (required) option
-logic {AND|OR}.And, just to state explicitly, the full probabilistic tracking in
-mode PROBdoes not (currently) produce tract file output. Such is life and also an impetus behind the mini-probabilistic methodology (described further below).
7.5.5. Viewing tracked outputs¶
Many different types of output files can be viewed simultaneously in SUMA (volume, tractfile, dset/matrices, etc.). SUMA and AFNI can also be run at the same time to talk together and share informative gossip on data sets. All the individual SUMA examples below can be combined in a single command line call. After opening a controller, you can hit the new useful ‘All Objs.’ button near the top, in order to immediately be able to toggle among each input file. For more information on SUMA viewing in general, check out SUMA Viewer.
Volume files outputs. PAIRMAP, INDIMAP and dumped volumes can all be viewed in either AFNI or in SUMA. To load them into the latter for 3D visualization, use:
suma -vol FILENAME ...
By default, they are displayed as slices and not as surfaces, but you can select that capability (see description in Volume Viewing).
To view the volume files in the 2D afni slice viewer, one uses the standard, general call to open AFNI (assuming you’re in a directory where those files are located; otherwise, include the path to them):
afniMatrix file outputs. SUMA is used to view the matrix information in the
*.dsetfile. While one can view this information as a ‘classic’ connectivity matrix (for both3dTrackIDand3dNetCorroutputs), it is also possible to view the data as coloration of graph edges and/or tract bundles in the brain volume. For more features, please see the help examples in SUMA: Graph Viewing. To load the data into SUMA, use:suma -gdset FILE.niml.dset ...
Additionally, one can select, view and save the matrices from the command line with a Python-based tool,
fat_mat_sel.py. This program can output several matrices from several subjects simultaneously, and the user can control several features of the plotting (font size, colorbar properties, ranges, DPI, etc.). It can be useful, for example, when making outputs for presentations and publications. See the helpfile:fat_mat_sel.py -h
for more information and list of the options.
Tract files. These are viewable in SUMA with many, many interactive features. To load in the tracts:
suma -tract FILE.niml.tract ...
Default coloration is by local tract orientation, but one can also color, for example, by bundle (useful for connectomes) or by the connectivity matrix information (importing the
-gdset FILE.niml.dsetinformation, above).Selection masks (either sphere or box) can be made for specifying subsets of tracts. One can have multiple selection masks, and use AND- and/or OR-logic with them. Importantly, these volumes are dragged along the tracts and bundles themselves, so that one can follow arbitrary trajectories through 3D (i.e., one is not constrained to manipulating them just in 2D slices).
For more information, please see the voluminous set of features, hints and examples in the SUMA help: Tract Viewing.
TRK files. These
NAME.trkfiles are generated using the TrackVis format, and as such can be viewed in the eponymous program. (They are not output by default.)
7.5.6. Inputs for tracking¶
This section will be an attempt to cluster 3dTrackID input options meaningfully.
Minimal inputs for each mode¶
Each option is briefly explained the first time it is mentioned; one
can assume that, unless explicitly noted, the initial definition still
holds. A selection of -mode {DET|MINIP|PROB} is always required,
as well.
The examples are shown for DTI tracking, and the simple option change in each case for performing HARDI tracking is provided immediately after.
Deterministic (DET) DTI:
3dTrackID -mode DET \ -dti_in DT_PREF \ -netrois TARGET_ROI_FILE \ -logic {AND|OR} \ -prefix OUT_PREF
where:
-dti_in DT_PREF: point to the set of DTI parameter files by their prefix. The program will read in all scalar files with this prefix and output WM ROI statistics on them. The minimum set of files needed for tracking is:(scalar) FA, MD and L1– RD is calculated automatically if it’s not loaded in
(vector) V1, V2 and V3
The function will glob for all scalar files with the entered prefix (
-dti_in DT_PREFleads to searching for file names like ‘DT_PREF*’), so other scalars can be easily included for automatic connectivity matrix calculation by giving them the same prefix. (See below for other ways of including extra files.)-netrois TARGET_ROI_FILE: input the file of targets among which to find connections. This can be a file with multiple volumes/bricks, and each brick is treated like a separate network. Each target in a network is defined as a set of voxels with a given integer, and a labletable can be attached for further target naming with strings (with the labels also being attached to tracked outputs).-logic {AND|OR}: select whether the tracts output in the *.tract file connect targets using AND- or OR-logic. NB: in either case, both INDI and PAIR map (volume) files are output.-prefix OUT_PREF: prefix for all output files, as described above. Additionally, a network number will be appended before the file extensions, starting with 000, 001, 002, etc. (in order to match the brick number of the-netroisfile).
Note
Instead of
-dti_in DT_PREF, one can input an explicit file of list of DTI parameter files to input in a niml-formatted text file with-dti_list FILE.niml.opts. An example is provided in the 3dTrackID help file under “DTI LIST FILE EXAMPLE”. Up to 4 ‘extra’ scalar-valued files can be input for statistical purposes.Mini-probabilistic (MINIP) DTI:
3dTrackID -mode MINIP \ -dti_in DT_PREF \ -netrois TARGET_ROI_FILE \ -logic {AND|OR} \ -uncert U_FILE \ -mini_num NREP \ -prefix OUT_PREF
where:
-uncert UNCERT_FILE: the file of uncertainty values output by 3dDWUncert.-mini_num NREP: the number of perturbed Monte Carlo repetitions to perform. Often 5-7 seems to be a good number.
Fully probabilistic (PROB) DTI:
3dTrackID -mode PROB \ -dti_in DT_PREF \ -netrois TARGET_ROI_FILE \ -uncert U_FILE \ -prefix OUT_PREF
where: all the options have been described in the previous two examples! (This method produces no tract results, however, just volumes. But those can be quite useful, too.)
Performing HARDI tracking in each of the above cases is done with a change of one option:
Replace
-dti_in DT_PREF(or-dti_list FILE.niml.opts) with:-hardi_gfa GFA: the scalar map which you want to be thresholded to contrain the propagation (that is, a generalization of what the FA map typically does for DTI tracking);-hardi_dirs DIRS: the file of vectors in X>1 directions. The assumed format of FILE2 is to have 3*X bricks of (x1, y1, z1, x2, y2, z2, …) ordered, unit magnitude vector components.
Also, note that when using HARDI data for either MINIP or PROB tracking, then the uncertainty file must have a different format than the one output by 3dDWUncert for DTI tracking. It must have X+1 briks, where U_FILE[0] is the uncertainty for the GFA (scalar) file, and the other briks are ordered for directions given with the DIRS file (vectors; uncertainty in this case is characterized by a single angle, sweeping out a cone of uncertainty).
Including extra volumes¶
One might want to load extra volumes of information into
3dTrackIDfor making extra connectivity matrices in the output *.grid files. For example, one might want statistics performed on non-diffusion data such as T1 or PD values.If using
-dti_in DT_PREF, one can give these files the same prefix, so that they are found using the glob for ‘DT_PREF*’ filenames.If using
-dti_list FILE.niml.opts, one can enter the other filenames directly (without special prefix), in the NIML-formatted file; see the second example under “DTI LIST FILE EXAMPLE” in the 3dTrackID help.For the HARDI data case, one can input a prefix using
-hardi_pars PREFand glob for all single brick files with the name ‘PREF*’.
Alternatively, in DTI analysis one might want to use a non-FA map to restrict tract propagation, for example using a T1-weighted segmentation. For this purpose, one would load it in using
-dti_extra SET. In grid files, name of this quantity will be ‘XF’ (stands for ‘extra file’).NB: if the file
SEThappens to have a name like ‘DT_PREF*’, it will still be globbed for using-dti_in DT_PREF, and therefore included twice. But that shouldn’t harm any results.Note
To turn off the globbing capability (beyond finding just the bare minimum DTI files), one can use the
-dti_search_NOswitch.
Including tract masks¶
One can restrict all tracts to lie within a mask using
-mask MASK. (If no MASK is input, then internally some automasking is performed; often, DTI has already been masked to include just the whole brain, which would then be used as the internal mask.)Alternatively, if you want to allow tracts anywhere in the brain but to keep only those which pass through a particular region, then you can load that region in as a “thru-mask” with
-thru_mask TM.And, though it’s not a separate option, if you want to make an “anti-mask” region through which tracts are not allowed to go, you can give that region negative values in the particular network loaded in with
-netrois TARGET_ROI_FILE.
Changing default tracking parameters¶
The following major tracking parameters can all be changed individually from the command line (default values are given):
for all modes:
-alg_Thresh_FA A: set threshold for DTI FA map, ‘-dti_extra’ FILE, or HARDI GFA map (default = 0.2).-alg_Thresh_ANG B: set max angle (in deg) for turning when going to a new voxel during propagation (default = 60).-alg_Thresh_Len C: min physical length (in mm) of tracts to keep (default = 20).for
{DET|MINIP}modes:-alg_Nseed_X D: Number of seeds per vox in x-direc (default = 2).-alg_Nseed_Y E: Number of seeds per vox in y-direc (default = 2).-alg_Nseed_Z F: Number of seeds per vox in z-direc (default = 2).for
PROBmode:-alg_Thresh_Frac G: value for thresholding how many tracks must pass through a voxel for a given connection before it is included in the final WM-ROI of that connection. It is a decimal value <=1, which will multiply the number of ‘starting seeds’ per voxel, Nseed_Vox*Nmonte (see just below for those; default = 0.001; for higher specificity, a value of 0.01-0.05 would be used).-alg_Nseed_Vox H: number of seeds per voxel per Monte Carlo iteration; seeds will be placed randomly (default = 5).-alg_Nmonte I: number of Monte Carlo iterations (default = 1000).
The above
alg_*tracking parameters can also be set at once in a single text file. The text file can either have only plain text and no labels, or it can be in NIML-format with nice labels so that there’s no confusion about which value is being set. See the3dTrackIDhelp file’s “ALGOPT FILE EXAMPLES” for more information. The option file is loaded in using-algopt A_FILE.When in MINIP and PROB modes, which use the uncertainty of parameter values, one can choose an explicit minimum uncertainty; in general, the uncertainty files will have been generated using
3dDWUncert, but for whatever reason you might want to enforce a minimal angular uncertainty or something. The values are set with:-unc_min_FA VAL1: the minimum stdev for perturbing FA (in-dti_in), or the EXTRA- file also in DTI (-dti_extra), or GFA (in-hardi_*). Default value is: 0.015 for FA, and 0.015 times the max value in the EXTRA-file or in the GFA file.-unc_min_V VAL2: the minimum stdev for perturbing eigen-/direction-vectors. In DTI, this is for tipping V1 separately toward V2 and V3, and in HARDI, this is for defining a single degree of freedom uncertainty cone. Default values are 0.06 rad (~3.4 deg) for any eigenvector/direction. User assigns values in degrees.
Thresholding {DET|MINIP} bundles by tract count¶
The PROB method requires a certain number of tracts to go through a voxel before it is included in a WM ROI connection.
Recently, the ability to trim some kinds of ‘obvious’ noisy tracts
from DET and MINIP modes has been added. The option -bundle_thr V
allows the user to enter a minimum threshold number of tracts for
any bundle to have without being filtered out (AKA removed).
It is based on the fact that occasionally, one will see an odd tract winding as a connection between two targets, in what would appear visually to be an outlier. Even when using more DET seeds or MINIP iterations, the tract might remain isolated– further justifying its interpretation as noise-driven. The bundle threshold criterion can be useful in removing it easily.
Note, however, that the fully probabilistic mode’s criterion is stricter, and it still provides the most robust results when tracking.
Using target surfaces to control tract trimming¶
In each iteration of 3dTrackID, an initial set of all possible
tracts throughout the brain are generated, tracking forward and
backward as far as the stopping conditions allow from seeds in every
WM voxel (such as where FA>0.2). A network map is then “stamped”
down, and any tracts that intersect targets are kept (with the others
temporarily ignored). For connections between pairs of targets, once
can decide how much of the initial tract constitutes a connection.
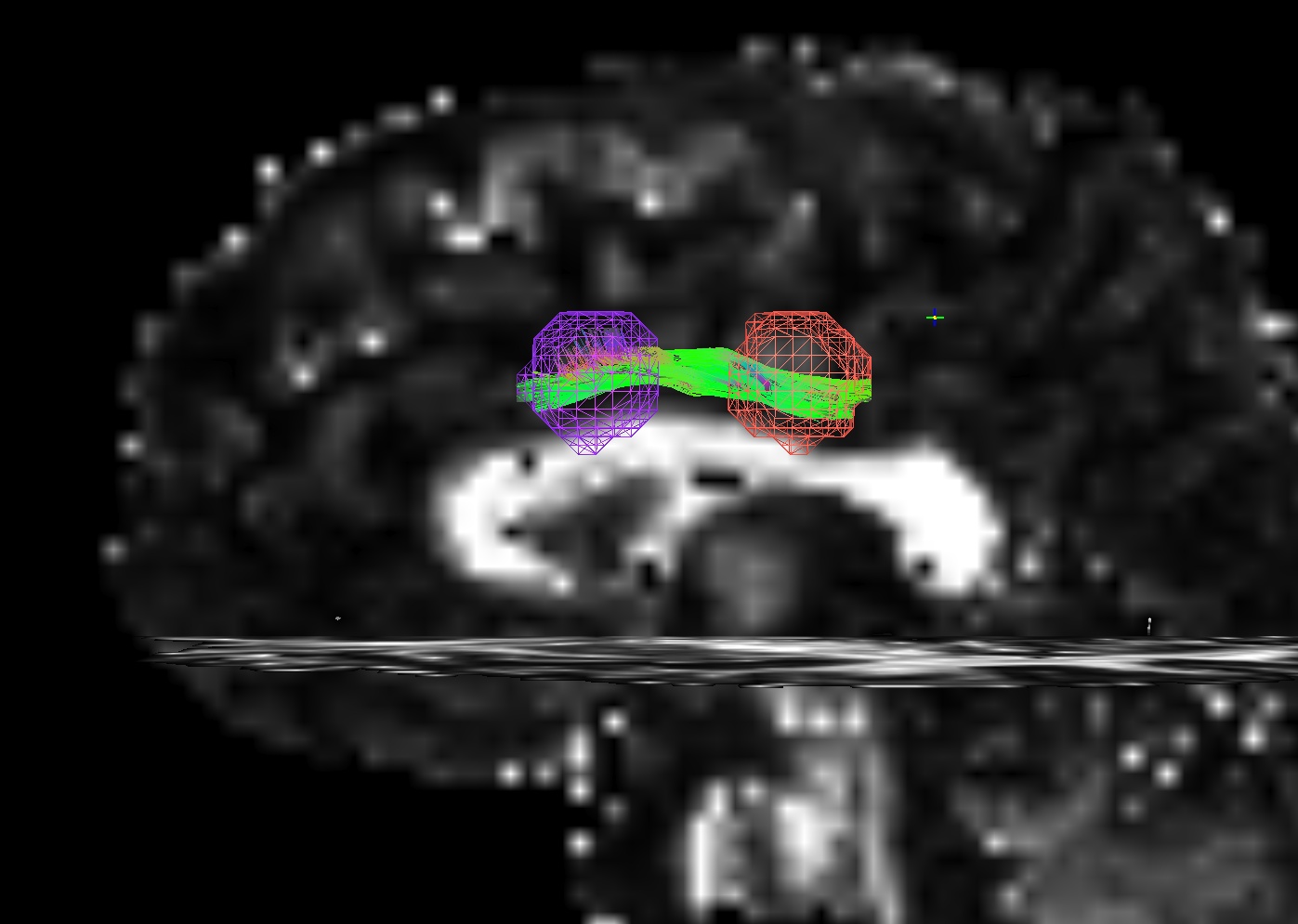
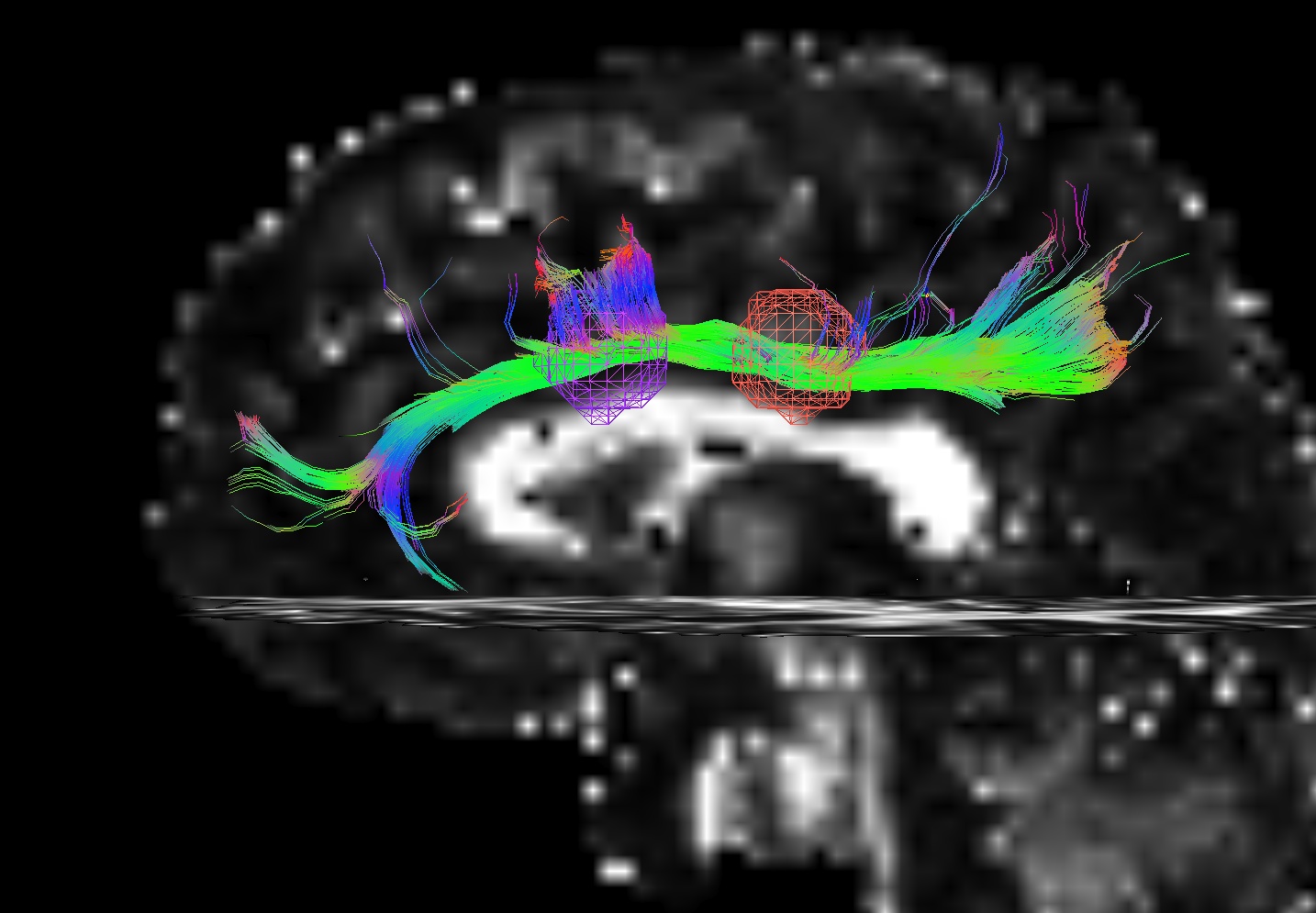
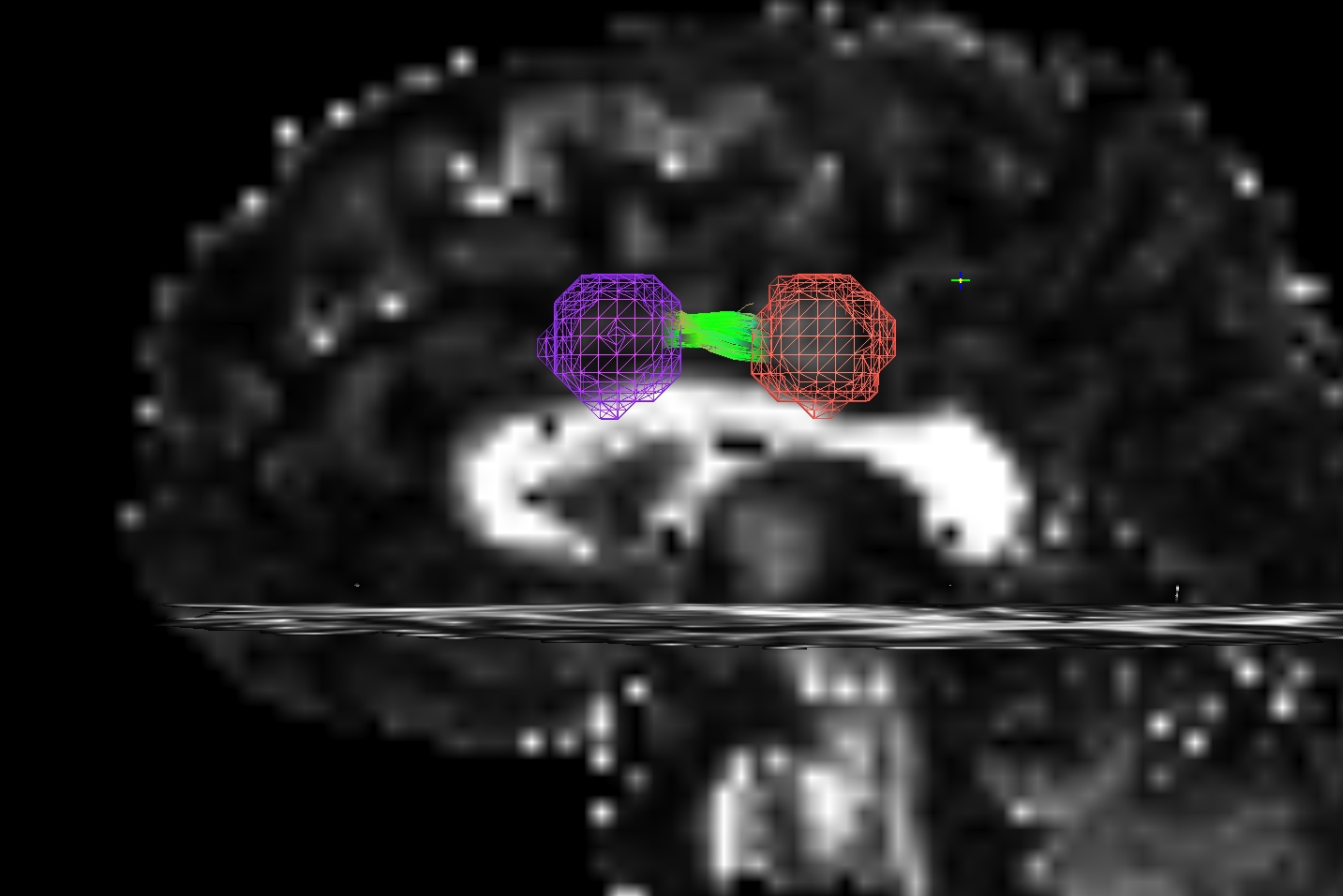
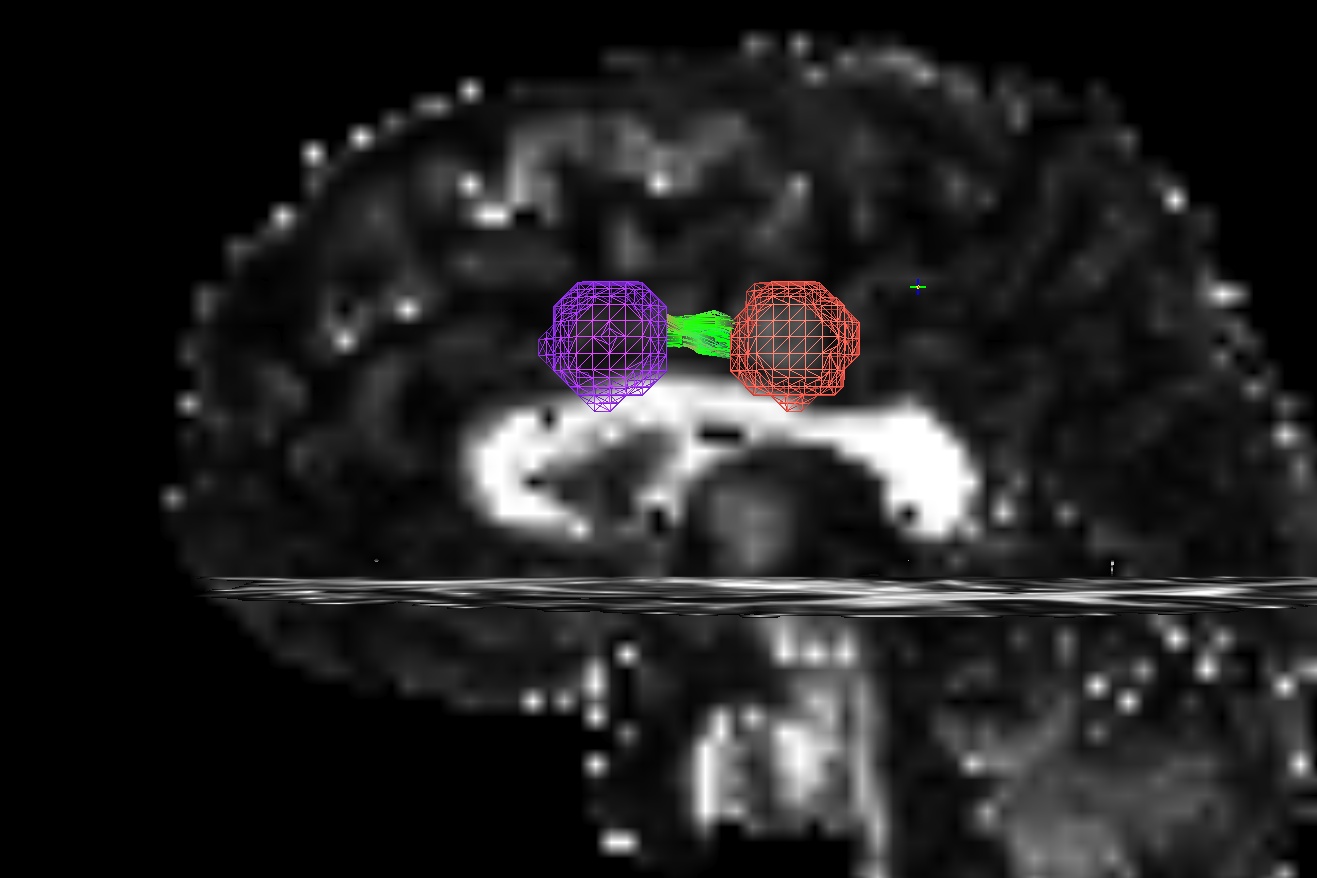
The figure below demonstrates the four current possibilities (AFNI version >=16.3.09). It could be:
Default: only the parts of the tract within and between the targets; that is, parts of the initial tract that stick out away from the partner target are ignored.
-uncut_at_rois: The whole initial tract: parts of the tract within each target, between the targets, and those endparts sticking out away from each target.-targ_surf_stop: only parts of the tract between the targets and just one layer into the target volumes; that is, the target surface stops the tracts after they enter.-targ_surf_twixt: only parts of the tract between the targets, stopping just just outside of the target volumes; that is, the tracts are only between (= betwixt) the targets, not overlapping at all.
Each of these approaches applies to any mode of tracking (DET,
MINIP or PROB). Also, each approach only refers to the
AND-logic (= pairwise; the off-diagonal elements of the returned
matrix of structural properties) connections between two targets; the
OR-logic tracts (the on-diagonal elements in the matrix of structural
properties) that are defined by going through at least a single target
are unaffected (those are always untrimmed).
Note
Note that for the -targ_surf_* cases, a tract between
targets A and B could overlap/pass through a separate
target C, though; the restriction on a tract’s overlap only
refers to the two targets it connects. Separate tracts from
A to C and from B to C would obey specified surface-stopping
rules with those respective targets.
Tract control options in |
|
|---|---|
|
|
|
|
|
|
|
|
Using SUMA for visualization (sagittal view, FA slice as background), the above figure shows two targets (represented as magenta and orange meshes) and various ways that the AND-logic, pairwise connection tracts for them could be returned.
Useful output dumping of WM ROIs¶
See the -dump_rois * option above in Outputs common to all modes. I think
it’s pretty valuable to use one of -dump_rois {AFNI|AFNI_MAP}, in
order to be able to have individual WM ROI files output. The PAIR and
INDI maps are mostly for quick reference, in my opinion, while the
dumped files can be more useful in viewing or further quantitative
analyses.
Switching various features ON/OFF¶
-do_trk_out: do output *.trk files, which might be useful in other, non-AFNI/SUMA programs.-uncut_at_rois: by default, tracts connecting pairs of targets are restricted to lie within and between the targets– if a tract carries on through the other side, that part is cut and not recorded as part of the pair’s ‘connection’. If you don’t want this trimming process to occur, then use this switch.-no_indipair_out: choose to not output a PAIR and INDI map. Might be useful to save space if one has a lot of targets in a network. On could utilize this switch and then just use the-dump_rois *option, as well.-write_opts: output a NIML-formatted file of the algorithm options being used. Might be useful if you want to keep it around to use later or as a record.-write_rois: write out a file (PREFIX.roi.labs) of all the ROI (re-)labels, for example if the input ROIs aren’t simply consecutive and starting from 1. The file has three cols: Input_ROI, Condensed_form_ROI, Power_of_2_label.-dump_no_labtab: if the ROIS file has a label table, the default is to use it in naming a-dump_rois *output (if being used); using this switch turn that off– output file names will be the same as if no label table were present.
Miscellaneous others¶
-nifti: output all volume files as*.nii.gzfiles.-extra_tr_par: run three extra track parameter scalings for each target pair, output in the *.grid file. The NT value of each connection is scaled in the following manners for each subsequent matrix label:NTpTarVol: div. by average target volume;
NTpTarSA: div. by average target surface area;
NTpTarSAFA: div. by average target surface area bordering suprathreshold FA (or equivalent WM proxy definition).
NB: the volume and surface area numbers are given in terms of voxel counts and not using physical units (consistent: NT values themselves are just numbers.)
Sundry other options described in the
3dTrackIDhelpfile (which most likely aren’t interesting enough to describe further): ‘-dump_lab_consec’, ‘-posteriori’, ‘-rec_orig’ and ‘-pair_out_power’.
