12.5.6. Preproc: TORTOISE¶
DIFFPREP¶
This function is run differently depending on whether one plans to run
DR_BUDDI afterwards or not. Basically, if the DWIs have been
acquired as a dual phase encoded (AP-PA or “blip up/blip down”) data
set, then you will use DR_BUDDI for EPI distortion reduction; if
not, then you won’t. Here, we do have AP-PA data, and so use
DIFFPREP accordingly on each of the AP and PA sets. The final
outputs of interest will be a set of motion- and eddy-distortion
reduced set of DWIs, along with b-matrices that have been adjusted
for any rotations applied the volumes.
Proc: For pipeline simplicity and modularity, we make a separate
working directory for the DIFFPREP inputs, because some
reformatting is necessary, and this directory ($odir) will also
hold the processed files. Essentially the same steps are applied to
each of the AP and PA sets, creating two separate directories.
The main inputs (volume + gradient info) can be provided to
DIFFPREP in two different formats: either via a listfile, or as
command line arguments. We use the latter here, which means we need to
make separate row-wise files of the b-values and (unscaled)
b-vector gradients; this is accomplished with a single call to
1dDW_Grad_o_Mat++. The NIFTI volume of DWIs must be unzipped,
accomplished during the 3dcopy step. The reference anatomical
merely gets uncompressed (once, since the same volume is used in both
the AP and PA processing).
For DIFFPREP itself, most of the inputs are fairly
self-explanatory. Note that we flag that we will be following this
processing with DR_BUDDI (via --will_be_drbuddied 1). We also
note that the phase-encode direction was along the A-P axis of the
volume (that is what --phase vertical signifies). Finally, we just
use the default settings file from the TORTOISE folks (via
--reg_settings *.dmc; note that this text file should be sitting
in a directory with the following location and name:
$HOME/DIFF_PREP_WORK/). For the AP data set, we then execute:
# I/O path, same as above, following earlier steps
set path_P_ss = data_proc/SUBJ_001
# make a directory to hold 'starter' data for DIFFPREP, as well
# as all the files it creates
set odir = "$path_P_ss/dwi_03_ap"
if ( ! -e $odir ) then
mkdir $odir
endif
# uncompress the anatomical
gunzip $path_P_ss/anat_01/t2w.nii.gz
# for DIFFPREP command line, need row-vec and row-bval format
1dDW_Grad_o_Mat++ \
-in_col_matT $path_P_ss/dwi_02/ap_matT.dat \
-unit_mag_out \
-out_row_vec $odir/ap_rvec.dat \
-out_row_bval_sep $odir/ap_bval.dat
# the NIFTI file must be unzipped
3dcopy \
$path_P_ss/dwi_02/ap.nii.gz \
$odir/ap.nii
# finally, the main command itself
DIFFPREP \
--dwi $odir/ap.nii \
--bvecs $odir/ap_rvec.dat \
--bvals $odir/ap_bval.dat \
--structural $path_P_ss/anat_01/t2w.nii \
--phase vertical \
--will_be_drbuddied 1 \
--reg_settings TORTOISE_AFNI_bootcamp_DATA_registration_settings.dmc
... and for the PA data set, nearly the same command with “ap” -> “pa”:
# I/O path, same as above, following earlier steps
set path_P_ss = data_proc/SUBJ_001
# make a directory to hold 'starter' data for DIFFPREP, as well
# as all the files it creates
set odir = "$path_P_ss/dwi_03_pa"
if ( ! -e $odir ) then
mkdir $odir
endif
# for DIFFPREP command line, need row-vec and row-bval format
1dDW_Grad_o_Mat++ \
-in_col_matT $path_P_ss/dwi_02/pa_matT.dat \
-unit_mag_out \
-out_row_vec $odir/pa_rvec.dat \
-out_row_bval_sep $odir/pa_bval.dat
# the NIFTI file must be unzipped
3dcopy \
$path_P_ss/dwi_02/pa.nii.gz \
$odir/pa.nii
# finally, the main command itself
DIFFPREP \
--dwi $odir/pa.nii \
--bvecs $odir/pa_rvec.dat \
--bvals $odir/pa_bval.dat \
--structural $path_P_ss/anat_01/t2w.nii \
--phase vertical \
--will_be_drbuddied 1 \
--reg_settings TORTOISE_AFNI_bootcamp_DATA_registration_settings.dmc
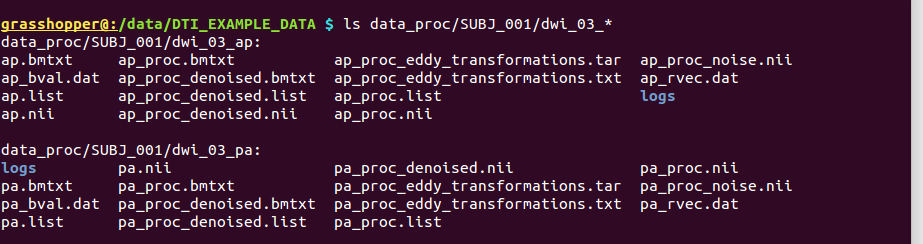
-> producing separate subdirectories ‘data_proc/SUBJ_001/dwi_03_ap/’ and ‘data_proc/SUBJ_001/dwi_03_pa/’:
Directory substructure for example data set |
|---|
|
Output subdirectories made by TORTOISE’s DIFFPREP applied separately to the AP and PA data sets. |
In the present set of steps, these outputs are intermediate to running
DR_BUDDI, and so we continue on. Note that it would be good to
look at these outputs.
DR_BUDDI¶
Proc. This step combines both the AP and PA data sets; it makes
use of the transforms calculated earlier, and will combine each with
subsequent ones calculated here and then apply them. Therefore, the
“list” files of the data processed by DIFFPREP are provided. The
same anatomical reference volume is also used.
run:
# I/O path, same as above, following earlier steps
set path_P_ss = data_proc/SUBJ_001
DR_BUDDI_withoutGUI \
--up_data $path_P_ss/dwi_03_ap/ap_proc.list \
--down_data $path_P_ss/dwi_03_pa/pa_proc.list \
--structural $path_P_ss/anat_01/t2w.nii \
--distortion_level medium \
--res 1.5 1.5 1.5 \
--output $path_P_ss/dwi_04/buddi.list
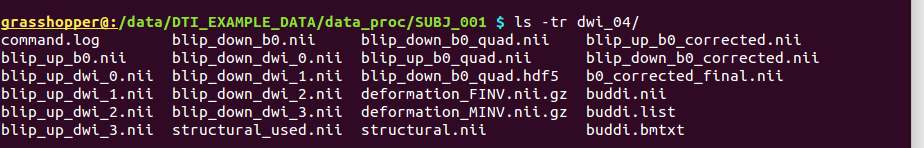
-> producing a single subdirectory ‘data_proc/SUBJ_001/dwi_04/’
(though some files are also added to the locations of the input
*_proc.list files):
Directory substructure for example data set |
|---|
|
Output subdirectory made by TORTOISE’s DR_BUDDI applied to the results of DIFF_PREP, combining the AP and PA sets. |
It contains the following outputs of both DWI and anatomical data
sets. Note that the volumes can be grouped into two sets by space, 1)
ones that are in the “blip_up ” space (which, for our
purposes, are all early/intermediate files), and 2) ones that are in
the final “structural” registration space (which are ones that we will
use). Further note that some of the names are familiar, so
doublecheck to make sure that you are using the correct volume in your
scripts.
Outputs of |
TORTOISE’s |
|---|---|
command.log |
textfile, copy of the command that was run. |
blip*.nii, deformation*nii.gz, b0_corrected_final.nii |
some intermediate volumes in “blip_up |
structural_used.nii |
volumetric NIFTI file, 3D (single brick volume), the reference
anatomical in the same resolution as it was input, but in
“blip_up |
structural.nii |
volumetric NIFTI file, 3D (single brick volume), the reference anatomical in “structural” space, which will define the spatial resolution+grid as the output DWIs (i.e., it is probably resampled from its own original input); not to be confused with structural_used.nii, above. |
buddi.nii |
volumetric NIFTI file in “structural” space, 4D (N=31 volumes), the final DWI volumes (AP and PA information combined into one), in which we are highly interested. |
buddi.bmtxt |
textfile, column file of (DW scaled) TORTOISE-style b-matrix
( |
buddi.list |
textfile, list of names of final output DWI volume and b-matrix files, as well as the phase encode direction. |
Note
In general (at least for TORTOISE v3.0), the origin and
orientation of the FOVs of these volumes will be different
than what was input. Therefore, don’t expect other data
sets to overlay on these immediately; making these volumes
play more nicely with the other data sets is part of the
role of later fat_proc commands.

Comments¶
This stage describes performing the “actual” preprocessing of the DWIs themselves, in terms of reducing the effects of distortions due to subject motion, eddy currents and B0 inhomogeneities. This collective action is usually called “distortion correction,” although we should be clear that at best we can only reduce effects of distortions retrospectively. The degree to which that desirable goal is possible depends on the processing tools being used, but also (heavily) on the study design and on the acquired data. In practice, as well, the practical effectiveness of processing also depends on the type of analysis to be performed afterwards, because the answer to the question, “Are my data clean enough to be appropriate for analysis X?” depends on what the details and assumptions of that testing are.
These major steps are performed using the TORTOISE tools, which are also freely available from the NIH (thanks, taxpayers!). Here, we describe briefly how we use
DIFFPREPandDR_BUDDIin TORTOISE v3.* (no description ofDIFF_CALC; we perform fitting with AFNI-FATCAT tools).The data set used here has the following characteristics:
pretty clean, not super distorted or noisy
acquired on a 3T scanner
from a human
from an (ostensibly healthy) adult.
:math:`rightarrow` If your data are not so, then you should check with the TORTOISE folks about options and flags to use in the commands. Even if your data do match these characteristics, you should probably be in touch with them about finer points of this processing analysis. Here, we try to work closely with the TORTOISE folks, to keep in touch with the latest-and-greatest updates and to represent those here, but it is worth being veeery clear about fine details in processing.
Note
Disclaimer: while we work closely with the TORTOISE folks, trying to keep in touch with the latest-and-greatest updates and to represent those here, we are not full experts in it– and it is worth being veeery clear about fine details in processing. All TORTOISE-related questions about options or problems should be addressed to the FS gurus themselves. Though, we are also happy to be involved in discussions, and any feedback on things to do differently would be welcomed and gladly discussed on our end.
As noted previously, the TORTOISE folks have kindly provided valuable feedback and input here, so we appreciate them for that.
Finally, we note that there are some differences what we do here and the official TORTOISE gurus’ recommendations for the tools. For example, they recommend loading DICOMs directly into TORTOISE, to deal with header information internally; if weird things start happening in your data, please consider this. (Most importantly, again– please keep looking at your data to know what is happening at each step along the way!)
A comment on running earlier versions of TORTOISE. In the Dark Ages TORTOISE either required an IDL license to run in batchable mode or was run subject-by-subject by clicking through an IDL VM GUI (ick). A PDF describing a system of steps for processing with TORTOISE v2.5.2 is provided here, “
Running_TORTOISE_v2.5.2.pdf,” for the time being, in part for the nostalgia of youth. Earlier versions of TORTOISE (v2.5.1 and previous) have greater subtlety in processing, and are entirely ignored. We don’t comment any further about this version of processing here.